A fundamental goal of developmental biology is to understand the lineage relationships of cells and cell types to one another, as well as the molecular programs that underlie each cell type’s emergence. In principle, developmental programs can be comprehensively described, as in Sulston and colleagues’ heroic reconstruction of the complete embryonic lineage of the roundworm Caenorhabditis elegans1. However, C. elegans—small, translucent, and developmentally invariant—remains the only model organism for which such a complete description has been realized.
Since 2016, we and others have developed and applied new technologies for single-cell molecular profiling at the ‘whole-animal’ scale, including worm, fly, zebrafish, frog and mouse2,3,4,5,6,7. Such studies lay the foundations for global views of animal development, such as by populating the Sulston lineage of C. elegans with the gene expression programs of each cell type7,8.
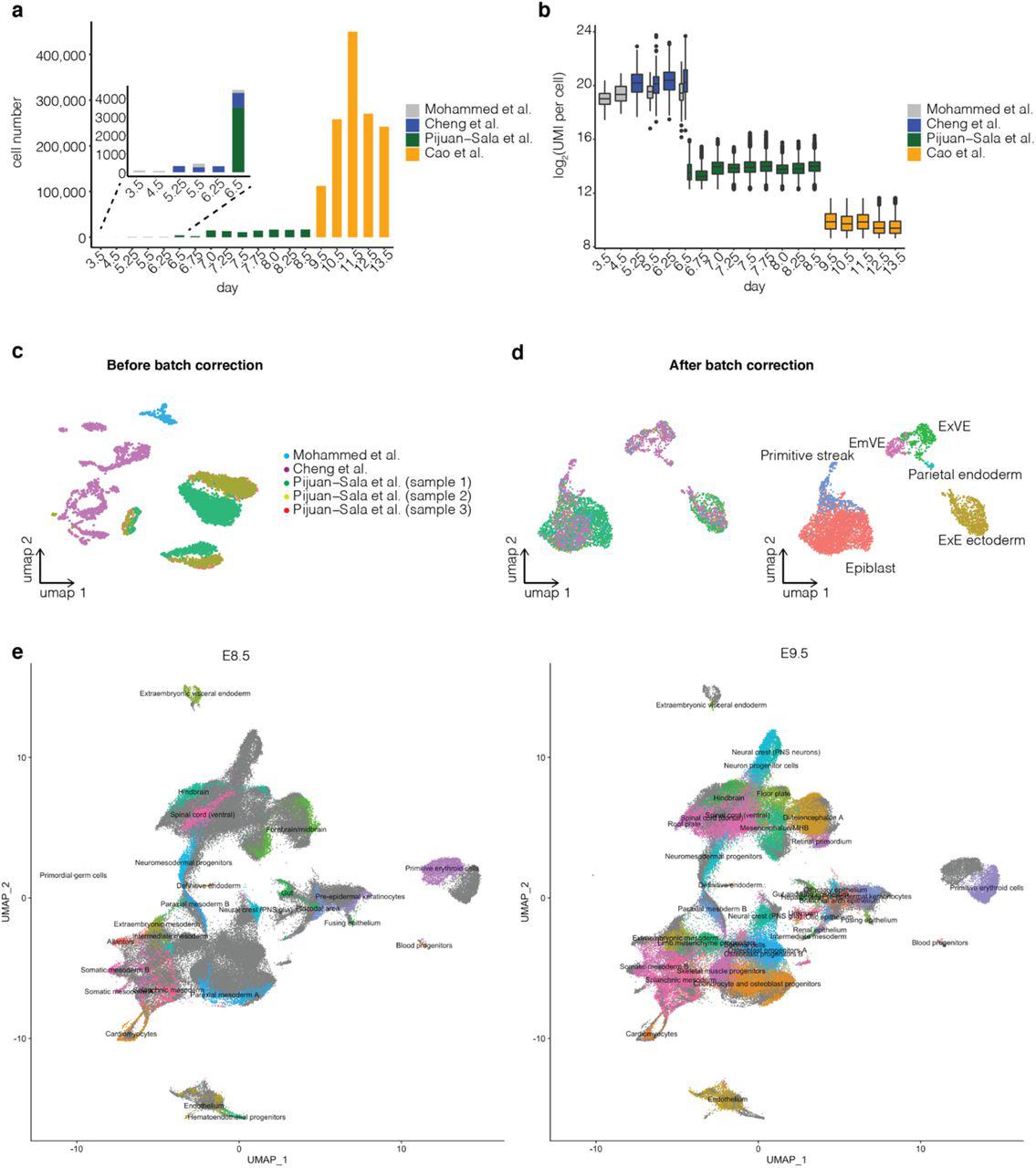
For mouse, the whole embryo has been profiled by scRNA-seq during implantation9,10, gastrulation2 and organogenesis4. Collectively, these studies span development from dozens of cells of a few types (E3.5) to millions of cells of hundreds of types (E13.5). However, the associated data have yet to be systematically integrated in a manner that permits their robust exploration. Such integration is challenging, both for technical reasons (e.g., different technologies and batch effects) and because of the sheer complexity of mouse development.
Here, we set out to systematically reconstruct the major cellular trajectories of mammalian embryogenesis from E3.5 to E13.5. Our primary strategy, inspired by Briggs and colleagues5, makes several assumptions: (1) although mouse development is variable, key patterns will be consistent across animals; (2) omnis cellula e cellula also applies to cell types (i.e., cell types observed at a given time point must have arisen from cell types present at the preceding time point); (3) we are sampling frequently and deeply enough that newly detected cell types will not arise from antecedent cell types undetected at the preceding time point; and (4) assuming sampling time points are closely spaced, transcriptional similarity is an effective means of linking related cell types across time.
A caution is that in contrast to the Sulston et al.’s seminal map of C. elegans, we focus here on reconstructing trajectories11, a concept related, but by no means equivalent, to lineage. Although it is a reasonable expectation that closely related cells (e.g., siblings) will be transcriptionally similar8, the converse is not necessarily true. For example, lineally distant cells might be insufficiently divergent, or even convergent, obscuring lineage relationships12. Furthermore, even the expectation that closely related cells will be transcriptionally similar is not always met, as rapid changes can lead to ‘gaps’ in trajectories8. In sum, our goal here is a continuous, navigable roadmap of the transcriptional states of cell types during mouse development. Such a roadmap may constrain the potential lineage relationships among constituent cell types, but it does not explicitly specify them.